The KBS Molecular Ecology and Genomics (MEG) Lab is a state of the art research and teaching laboratory established in 2017 to support the growing research and teaching conducted at KBS that involves molecular tools and bioinformatic analyses. This shared facility supports a diverse set of Next-Generation-Sequencing (NGS) pipelines, from DNA/RNA extraction through library preparation and sample send-off. The ultimate goal is to advance research that couples ‘omics’ studies with field based natural history and long-term fields studies.
The MEG lab is widely used by resident and visiting faculty, students, and research associates. Contact Sarah Fitzpatrick for more information.
Major equipment
The MEG lab is equipped with the full range of molecular research equipment necessary for DNA and RNA extraction, PCR, gel electrophoresis, and various genomic library preparation protocols. We have an Apple iMac dedicated to bioinformatics and working with molecular data and have access to Michigan State University’s High Performance Computing Cluster. In addition to standard equipment (centrifuges, -20 and -80 freezers, pipettes, thermocyclers) we have the following specialized equipment available for use:
- Agilent 2100 Bioanalyzer
- NuAire Class II, Type B2 Biological safety cabinet
- Covaris M220 Focused-ultrasonicator
- Eppendorf liquid handling robot
- Kingfisher Flex Purification System
- Milli-Q Integral water purification system
- Nanodrop 2000
- Qubit3 fluorometer
- Qiagen Tissue Lyser
- SmithRoot ANDe eDNA backpack sampler
Conducting research at the MEG Lab
Visiting scholars as well as resident graduate students, postdocs, and faculty use the MEG Lab for research. All use of this facility requires prior approval from the MEG Lab Coordinator, Sarah Fitzpatrick. If interested in conducting research in the MEG Lab, please see Conducting Research at KBS for more information and complete a KBS Site Use Request form. A new KBS Site Use Request (SURF) form must be submitted each calendar year prior to starting work on a project.
Fee structure for MEG lab use
| Small Project | Medium Project | Large Project | |
|---|---|---|---|
| User fee | $750 | $1500 | $3000 |
| Lab Use (in hours) | 5 - 80 | 81 - 360 | 361 + |
Project size will be estimated at start of project. Final fees will be determined by working hours in the lab. The maximum charge for use of the MEG Lab per year will be $3000. NOTE: All MEG Lab users must complete an orientation prior to starting project. Orientation hours do not count towards billable project hours.
Research highlights
Example projects:
Fitness effects of loss-of-function mutations in duplicate genes – Conner Lab in collaboration with Shin-Han Shiu (MSU) and Patrick Krysan (Wisconsin) ; funded by NSF
Origins and adaptations of weedy and crop radish – Conner Lab in collaboration with Kevin Childs and Patrick Edger (MSU); funded by Rackham Foundation
Microbial ecology and litter decomposition in a hyperarid desert – Evans lab in collaboration with Kathy and Peter Jacobson (Grinnell College); funded by Michigan State University and National Geographic Society.
Connecting nitrogen transformations mediated by the rhizosphere microbiome to perennial cropping system productivity in marginal lands. Evans Lab in collaboration with Lisa Tiemann, Jim Cole (MSU), Maren Friesen (Washington State University); funded by Department of Energy
Range-wide landscape genomics of a threatened species of Great Plains fish – Fitzpatrick Lab; funded by US Fish & Wildlife and Kansas Dept. of Parks & Tourism
Determining QTL underlying behavioral variation in Trinidadian guppies – Fitzpatrick Lab in collaboration with Kim Hughes (FSU) & Kim Hoke (CSU); funded by NSF
Featured publications:
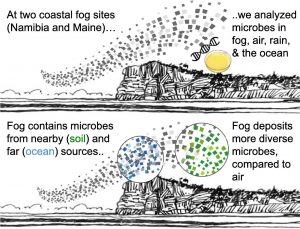
Evans, SE, ME Dueker, JR Logan, KC Weathers (2019) The biology of fog: results from coastal Maine and Namib Desert reveal common drivers of fog microbial composition. Science of The Total Environment 647:1547-1556 Link
Fitzpatrick, SW, BN Reid (2019) Does gene flow aggravate or alleviate maladaptation to environmental stress in small populations? Evolutionary Applications. in press Link



 .
. 



A legacy of conservation; a commitment to sustainability.
3700 E. Gull Lake Drive
Hickory Corners, MI 49060
(269) 671-5117
info@kbs.msu.edu